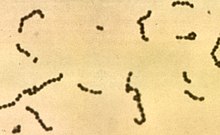Streptococcus
Streptococcus is a genus of gram-positive or spherical bacteria that belongs to the family Streptococcaceae, within the order Lactobacillales (lactic acid bacteria), in the phylum Bacillota.[2] Cell division in streptococci occurs along a single axis, thus when growing they tend to form pairs or chains, which may appear bent or twisted. This differs from staphylococci, which divide along multiple axes, thereby generating irregular, grape-like clusters of cells. Most streptococci are oxidase-negative and catalase-negative, and many are facultative anaerobes (capable of growth both aerobically and anaerobically).
The term was coined in 1877 by Viennese surgeon Albert Theodor Billroth (1829–1894),[3] by combining the prefix "strepto-" (from Ancient Greek: στρεπτός, romanized: streptós, lit. 'easily twisted, pliant'[4]), together with the suffix "-coccus" (from Modern Latin: coccus, from Ancient Greek: κόκκος, romanized: kókkos, lit. 'grain, seed, berry'.[5]) In 1984, many bacteria formerly grouped in the genus Streptococcus were separated out into the genera Enterococcus and Lactococcus.[6] Currently, over 50 species are recognised in this genus. This genus has been found to be part of the salivary microbiome.[7]
Pathogenesis and classification
[edit]In addition to streptococcal pharyngitis (strep throat), certain Streptococcus species are responsible for many cases of pink eye,[8] meningitis, bacterial pneumonia, endocarditis, erysipelas, and necrotizing fasciitis (the 'flesh-eating' bacterial infections). However, many streptococcal species are not pathogenic, and form part of the commensal human microbiota of the mouth, skin, intestine, and upper respiratory tract. Streptococci are also a necessary ingredient in producing Emmentaler ("Swiss") cheese.[9]
Species of streptococci are classified based on their hemolytic properties.[10] Alpha-hemolytic species cause oxidization of iron in hemoglobin molecules within red blood cells, giving it a greenish color on blood agar. Beta-hemolytic species cause complete rupture of red blood cells. On blood agar, this appears as wide areas clear of blood cells surrounding bacterial colonies. Gamma-hemolytic species cause no hemolysis.[11]
Beta-hemolytic streptococci are further classified by Lancefield grouping, a serotype classification (that is, describing specific carbohydrates present on the bacterial cell wall).[6] The 21 described serotypes are named Lancefield groups A to W (excluding E, I and J). This system of classification was developed by Rebecca Lancefield, a scientist at Rockefeller University.[12]
In the medical setting, the most important groups are the alpha-hemolytic streptococci S. pneumoniae and Streptococcus viridans groups, and the beta-hemolytic streptococci of Lancefield groups A and B (also known as "group A strep" and "group B strep").
Table: Medically relevant streptococci[10]
| Species | Host | Disease |
|---|---|---|
| S. pyogenes | human | pharyngitis, cellulitis, erysipelas |
| S. agalactiae | human, cattle | neonatal meningitis and sepsis |
| S. dysgalactiae | human, animals | endocarditis, bacteremia, pneumonia, meningitis, respiratory infections |
| S. gallolyticus | human, animals | biliary or urinary tract infections, endocarditis |
| S. anginosus | human, animals | subcutaneous/organ abscesses, meningitis, respiratory infections |
| S. sanguinis | human | endocarditis, dental caries |
| S. suis | swine | meningitis |
| S. mitis | human | endocarditis |
| S. mutans | human | dental caries |
| S. pneumoniae | human | pneumonia |
Alpha-hemolytic
[edit]When alpha-hemolysis (α-hemolysis) is present, the agar under the colony will appear dark and greenish due to the conversion of hemoglobin to green biliverdin. Streptococcus pneumoniae and a group of oral streptococci (Streptococcus viridans or viridans streptococci) display alpha-hemolysis. Alpha-hemolysis is also termed incomplete hemolysis or partial hemolysis because the cell membranes of the red blood cells are left intact. This is also sometimes called green hemolysis because of the color change in the agar.[citation needed]
Pneumococci
[edit]- S. pneumoniae (sometimes called pneumococcus), is a leading cause of bacterial pneumonia and the occasional etiology of otitis media, sinusitis, meningitis, and peritonitis. Inflammation is thought to be the major cause of how pneumococci cause disease, hence the tendency of diagnoses associated with them to involve inflammation. They possess no Lancefield antigens.[2]
The viridans group: alpha-hemolytic
[edit]- The viridans streptococci are a large group of commensal bacteria that are either alpha-hemolytic, producing a green coloration on blood agar plates (hence the name "viridans", from Latin vĭrĭdis, green), or nonhemolytic. They possess no Lancefield antigens.[2]
Beta-hemolytic
[edit]Beta-hemolysis (β-hemolysis), sometimes called complete hemolysis, is a complete lysis of red cells in the media around and under the colonies: the area appears lightened (yellow) and transparent. Streptolysin, an exotoxin, is the enzyme produced by the bacteria which causes the complete lysis of red blood cells. There are two types of streptolysin: Streptolysin O (SLO) and streptolysin S (SLS). Streptolysin O is an oxygen-sensitive cytotoxin, secreted by most group A Streptococcus (GAS), and interacts with cholesterol in the membrane of eukaryotic cells (mainly red and white blood cells, macrophages, and platelets), and usually results in beta-hemolysis under the surface of blood agar. Streptolysin S is an oxygen-stable cytotoxin also produced by most GAS strains which results in clearing on the surface of blood agar. SLS affects immune cells, including polymorphonuclear leukocytes and lymphocytes, and is thought to prevent the host immune system from clearing infection. Streptococcus pyogenes, or GAS, displays beta hemolysis.
Some weakly beta-hemolytic species cause intense hemolysis when grown together with a strain of Staphylococcus. This is called the CAMP test. Streptococcus agalactiae displays this property. Clostridium perfringens can be identified presumptively with this test. Listeria monocytogenes is also positive on sheep's blood agar.

Group A
[edit]Group A S. pyogenes is the causative agent in a wide range of group A streptococcal infections (GAS). These infections may be noninvasive or invasive. The noninvasive infections tend to be more common and less severe. The most common of these infections include streptococcal pharyngitis (strep throat) and impetigo.[13] Scarlet fever is another example of Group A noninvasive infection.
The invasive infections caused by group A beta-hemolytic streptococci tend to be more severe and less common. This occurs when the bacterium is able to infect areas where it is not usually found, such as the blood and organs.[14] The diseases that may be caused include streptococcal toxic shock syndrome, necrotizing fasciitis, pneumonia, and bacteremia.[13] Globally, GAS has been estimated to cause more than 500,000 deaths every year, making it one of the world's leading pathogens.[13]
Additional complications may be caused by GAS, namely acute rheumatic fever and acute glomerulonephritis. Rheumatic fever, a disease that affects the joints, kidneys, and heart valves, is a consequence of untreated strep A infection caused not by the bacterium itself, but due to the antibodies created by the immune system to fight off the infection cross-reacting with other proteins in the body. This "cross-reaction" causes the body to essentially attack itself and leads to the damage above. A similar autoimmune mechanism initiated by Group A beta-hemolytic streptococcal (GABHS) infection is hypothesized to cause pediatric autoimmune neuropsychiatric disorders associated with streptococcal infections (PANDAS), wherein autoimmune antibodies affect the basal ganglia, causing rapid onset of psychiatric, motor, sleep, and other symptoms in pediatric patients.
GAS infection is generally diagnosed with a rapid strep test or by culture.
Group B
[edit]S. agalactiae, or group B streptococcus, GBS, causes pneumonia and meningitis in newborns and the elderly, with occasional systemic bacteremia. Importantly, Streptococcus agalactiae is the most common cause of meningitis in infants from one month to three months old. They can also colonize the intestines and the female reproductive tract, increasing the risk for premature rupture of membranes during pregnancy, and transmission of the organism to the infant. The American College of Obstetricians and Gynecologists, American Academy of Pediatrics, and the Centers for Disease Control recommend all pregnant women between 35 and 37 weeks gestation to be tested for GBS. Women who test positive should be given prophylactic antibiotics during labor, which will usually prevent transmission to the infant.[15] Group III polysaccharide vaccines have been proven effective in preventing the passing of GBS from mother to infant.[16]
The United Kingdom has chosen to adopt a risk factor-based protocol, rather than the culture-based protocol followed in the US.[17] Current guidelines state that if one or more of the following risk factors is present, then the woman should be treated with intrapartum antibiotics:
- GBS bacteriuria during this pregnancy
- History of GBS disease in a previous infant
- Intrapartum fever (≥38 °C)
- Preterm labour (<37 weeks)
- Prolonged rupture of membranes (>18 hours)
This protocol results in the administration of intrapartum antibiotics to 15–20% of pregnant women and the prevention of 65–70% of cases of early onset GBS sepsis.[18]
Group C
[edit]This group includes S. equi, which causes strangles in horses,[19] and S. zooepidemicus — S. equi is a clonal descendant or biovar of the ancestral S. zooepidemicus — which causes infections in several species of mammals, including cattle and horses. S. dysgalactiae subsp. dysgalactiae[20] is also a member of group C, beta-haemolytic streptococci that can cause pharyngitis and other pyogenic infections similar to group A streptococci. Group C streptococcal bacteria are considered zoonotic pathogens, meaning infection can be passed from animal to human.[21]
Group D (enterococci)
[edit]Many former group D streptococci have been reclassified and placed in the genus Enterococcus (including E. faecalis, E. faecium, E. durans, and E. avium).[22] For example, Streptococcus faecalis is now Enterococcus faecalis. E. faecalis is sometimes alpha-hemolytic and E. faecium is sometimes beta hemolytic.[23]
The remaining nonenterococcal group D strains include Streptococcus gallolyticus, Streptococcus bovis, Streptococcus equinus and Streptococcus suis.
Nonhemolytic streptococci rarely cause illness. However, weakly hemolytic group D beta-hemolytic streptococci and Listeria monocytogenes (which is actually a gram-positive bacillus) should not be confused with nonhemolytic streptococci.
Group F streptococci
[edit]Group F streptococci were first described in 1934 by Long and Bliss among the "minute haemolytic streptococci".[24] They are also known as Streptococcus anginosus (according to the Lancefield classification system) or as members of the S. milleri group (according to the European system).
Group G streptococci
[edit]These streptococci are usually, but not exclusively, beta-hemolytic. Streptococcus dysgalactiae subsp. canis[20] is the predominant subspecies encountered. It is a particularly common GGS in humans, although it is typically found on animals. S. phocae is a GGS subspecies that has been found in marine mammals and marine fish species. In marine mammals it has been mainly associated with meningoencephalitis, sepsis, and endocarditis, but is also associated with many other pathologies. Its environmental reservoir and means of transmission in marine mammals is not well characterized. Group G streptococci are also considered zoonotic pathogens.
Group H streptococci
[edit]Group H streptococci cause infections in medium-sized canines. Group H streptococci rarely cause human illness unless a human has direct contact with the mouth of a canine. One of the most common ways this can be spread is human-to-canine, mouth-to-mouth contact. However, the canine may lick the human's hand and infection can be spread, as well.[25]
Clinical identification
[edit]
In clinical practice, the most common groups of Streptococcus can be distinguished by simple bench tests, such as the PYR test for group A streptococcus. There are also latex agglutination kits which can distinguish each of the main groups seen in clinical practice.
Treatment
[edit]Streptococcal infections can be treated with antibiotics from the penicillin family. Most commonly, penicillin or amoxicillin is used to treat strep infection. These antibiotics work by disrupting peptidoglycan production in the cell wall.[26] Treatment most often occurs as a 10-day oral antibiotic cycle. For patients with penicillin allergies and those suffering from skin infections, clindamycin can be used. Clindamycin works by disrupting protein synthesis within the cell.
Molecular taxonomy and phylogenetics
[edit]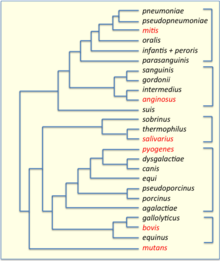
Streptococci have been divided into six groups on the basis of their 16S rDNA sequences: S. anginosus, S. gallolyticus, S. mitis, S. mutans, S. pyogenes and S. salivarius.[28] The 16S groups have been confirmed by whole genome sequencing (see figure). The important pathogens S. pneumoniae and S. pyogenes belong to the S. mitis and S. pyogenes groups, respectively,[29] while the causative agent of dental caries, Streptococcus mutans, is basal to the Streptococcus group.

Recent technological advances have resulted in an increase of available genome sequences for Streptococcus species, allowing for more robust and reliable phylogenetic and comparative genomic analyses to be conducted.[30] In 2018, the evolutionary relationships within Streptococcus was re-examined by Patel and Gupta through the analysis of comprehensive phylogenetic trees constructed based on four different datasets of proteins and the identification of 134 highly specific molecular signatures (in the form of conserved signature indels) that are exclusively shared by the entire genus or its distinct subclades.[30]
The results revealed the presence of two main clades at the highest level within Streptococcus, termed the "Mitis-Suis" and "Pyogenes-Equinus-Mutans" clades.[30] The "Mitis-Suis" main clade comprises the Suis subclade and the Mitis clade, which encompasses the Angiosus, Pneumoniae, Gordonii and Parasanguinis subclades. The second main clade, the "Pyogenes-Equinus-Mutans", includes the Pyogenes, Mutans, Salivarius, Equinus, Sobrinus, Halotolerans, Porci, Entericus and Orisratti subclades. In total, 14 distinct subclades have been identified within the genus Streptococcus, each supported by reliable branching patterns in phylogenetic trees and by the presence of multiple conserved signature indels in different proteins that are distinctive characteristics of the members of these 14 clades.[30] A summary diagram showing the overall relationships among the Streptococcus based on these studies is depicted in a figure on this page.
Genomics
[edit]
The genomes of hundreds of species have been sequenced.[32] Most Streptococcus genomes are 1.8 to 2.3 Mb in size and encode 1,700 to 2,300 proteins. Some important genomes are listed in the table.[33] The four species shown in the table (S. pyogenes, S. agalactiae, S. pneumoniae, and S. mutans) have an average pairwise protein sequence identity of about 70%.[33]
| feature | S. pyogenes | S. agalactiae | S. pneumoniae | S. mutans |
|---|---|---|---|---|
| base pairs | 1,852,442 | 2,211,488 | 2,160,837 | 2,030,921 |
| ORFs | 1792 | 2118 | 2236 | 1963 |
| prophages | yes | no | no | no |
Bacteriophage
[edit]Bacteriophages have been described for many species of Streptococcus. 18 prophages have been described in S. pneumoniae that range in size from 38 to 41 kb in size, encoding from 42 to 66 genes each.[34] Some of the first Streptococcus phages discovered were Dp-1[35][36] and ω1 (alias ω-1).[37][38][39] In 1981 the Cp (Complutense phage 1, officially Streptococcus virus Cp1, Picovirinae) family was discovered with Cp-1 as its first member.[40] Dp-1 and Cp-1 infect both S. pneumoniae and S. mitis.[41] However, the host ranges of most Streptococcus phages have not been investigated systematically.
Natural genetic transformation
[edit]Natural genetic transformation involves the transfer of DNA from one bacterium to another through the surrounding medium. Transformation is a complex process dependent on the expression of numerous genes. To be capable of transformation a bacterium must enter a special physiologic state referred to as competence. S. pneumoniae, S. mitis and S. oralis can become competent, and as a result actively acquire homologous DNA for transformation by a predatory fratricidal mechanism [42] This fratricidal mechanism mainly exploits non-competent siblings present in the same niche [43] Among highly competent isolates of S. pneumoniae, Li et al.[44] showed that nasal colonization fitness and virulence (lung infectivity) depend on an intact competence system. Competence may allow the streptococcal pathogen to use external homologous DNA for recombinational repair of DNA damages caused by the host's oxidative attack.[45]
See also
[edit]- Cia-dependent small RNAs
- Quellung reaction
- Streptococcal infection in poultry
- Streptococcal pharyngitis
- Streptokinase
References
[edit]- ^ a b c d e f g h i j k l m n o p q r s t u v w x y z aa ab ac ad ae af ag ah ai aj ak al am an ao ap aq ar as at au av aw ax ay az ba bb bc bd be bf bg bh bi bj bk bl bm bn bo bp bq br bs bt Parte AC. "Streptococcus". LPSN.
- ^ a b c Ryan KJ, Sherris JC, eds. (1994). Sherris Medical Microbiology (3rd ed.). Appleton & Lange. pp. 266–7. ISBN 0-8385-8541-8.
- ^ "streptococcus". Online Etymology Dictionary. Retrieved 25 July 2018.
- ^ στρεπτός in Liddell, Henry George; Scott, Robert (1940) A Greek–English Lexicon, revised and augmented throughout by Jones, Sir Henry Stuart, with the assistance of McKenzie, Roderick. Oxford: Clarendon Press. In the Perseus Digital Library, Tufts University.
- ^ κόκκος in Liddell and Scott
- ^ a b Facklam R (October 2002). "What happened to the streptococci: overview of taxonomic and nomenclature changes". Clinical Microbiology Reviews. 15 (4): 613–630. doi:10.1128/CMR.15.4.613-630.2002. PMC 126867. PMID 12364372.
- ^ Wang K, Lu W, Tu Q, Ge Y, He J, Zhou Y, et al. (March 2016). "Preliminary analysis of salivary microbiome and their potential roles in oral lichen planus". Scientific Reports. 6 (1): 22943. Bibcode:2016NatSR...622943W. doi:10.1038/srep22943. PMC 4785528. PMID 26961389.
- ^ "How to Get Rid of Pinkeye, Symptoms, Treatment, Causes & Pictures".
- ^ "Streptococcus | Center for Academic Research and Training in Anthropogeny (CARTA)". carta.anthropogeny.org. Retrieved 2022-07-23.
- ^ a b Patterson MJ (1996). Baron S; et al. (eds.). Streptococcus. In: Baron's Medical Microbiology (4th ed.). Univ of Texas Medical Branch. ISBN 978-0-9631172-1-2. (via NCBI Bookshelf).
- ^ Sharma S, Khanna G, Gangane SD (2019-07-13). Textbook of Pathology and Genetics for Nurses E-Book. Elsevier Health Sciences. ISBN 978-81-312-5538-4.
- ^ Carroll KC (August 2019). Munson E (ed.). "Biographical Feature: Rebecca Lancefield, Ph.D". Journal of Clinical Microbiology. 57 (8). doi:10.1128/JCM.00728-19. PMC 6663886. PMID 31142605.
- ^ a b c Cohen-Poradosu R, Kasper DL (October 2007). "Group A streptococcus epidemiology and vaccine implications". Clinical Infectious Diseases. 45 (7): 863–865. doi:10.1086/521263. PMID 17806050.
- ^ "Streptococcal Infections (Invasive Group A Strep)". New York City Department of Health and Mental Hygiene. Archived from the original on 6 November 2012. Retrieved 21 November 2012.
- ^ Schrag S, Gorwitz R, Fultz-Butts K, Schuchat A (August 2002). "Prevention of perinatal group B streptococcal disease. Revised guidelines from CDC". MMWR. Recommendations and Reports. 51 (RR-11): 1–22. PMID 12211284.
- ^ Noya, Francisco J. D.; Baker, Carol J. (1992-03-01). "PREVENTION OF GROUP B STREPTOCOCCAL INFECTION". Infectious Disease Clinics of North America. 6 (1): 41–55. doi:10.1016/S0891-5520(20)30424-4. ISSN 0891-5520. PMID 1578122.
- ^ "Prevention of Early-onset Neonatal Group B Streptococcal Disease: Green-top Guideline No. 36". BJOG. 124 (12): e280 – e305. November 2017. doi:10.1111/1471-0528.14821. PMID 28901693.
- ^ Norwitz ER, Schorge JO (2013). Obstetrics and Gynecology at a Glance (4th ed.). Chichester: John Wiley & Sons, Ltd. ISBN 978-1118341735.
- ^ Harrington DJ, Sutcliffe IC, Chanter N (April 2002). "The molecular basis of Streptococcus equi infection and disease". Microbes and Infection. 4 (4): 501–510. doi:10.1016/S1286-4579(02)01565-4. PMID 11932201.
- ^ a b Haslam DB, St Geme III JW (2023). "122 - Groups C and G Streptococci". In Long SS, Prober CG, Fischer M, Kimberlin D (eds.). Principles and Practice of Pediatric Infectious Diseases (Sixth ed.). Elsevier. pp. 752–753. doi:10.1016/B978-0-323-75608-2.00122-1. ISBN 978-0-323-75608-2. Note that according to the same source, the subspecies equisimilis is a grouping of large S. dysgalactiae colonies, whether they are members of Group C or Group G.
- ^ Klos, Marta (12 June 2017). "Pathogenicity of Virulent Species of Group C Streptococci in Human". Can J Infect Dis Med Microbiol. 2017: 1–5. doi:10.1155/2017/9509604. PMC 5485279. PMID 28694832.
- ^ Köhler W (June 2007). "The present state of species within the genera Streptococcus and Enterococcus". International Journal of Medical Microbiology. 297 (3): 133–150. doi:10.1016/j.ijmm.2006.11.008. PMID 17400023.
- ^ Holt et al. (1994). Bergey's Manual of Determinative Bacteriology (9th ed.). Lippincott Williams & Wilkins. ISBN 0-683-00603-7
- ^ Whitworth JM (November 1990). "Lancefield group F and related streptococci". Journal of Medical Microbiology. 33 (3): 135–151. doi:10.1099/00222615-33-3-135. PMID 2250284.[permanent dead link]
- ^ "Bacterial Infection (Streptococcus) in Dogs". petmd.com. Retrieved 12 December 2014.
- ^ Lowe, Derek (19 Jan 2022). "How Do Penicillins Actually Work?". Science – via American Association for the Advancement of Science.
- ^ "Bacteria-Firmicutes-Bacilli-Lactobacillales-Streptococcaceae-Streptococcus". PATRIC, University of Chicago. Retrieved 12 December 2014.
- ^ Kawamura Y, Hou XG, Sultana F, Miura H, Ezaki T (April 1995). "Determination of 16S rRNA sequences of Streptococcus mitis and Streptococcus gordonii and phylogenetic relationships among members of the genus Streptococcus". International Journal of Systematic Bacteriology. 45 (2): 406–408. doi:10.1099/00207713-45-2-406. PMID 7537076.
- ^ Liu, D., Molecular Detection of Human Bacterial Pathogens (Boca Raton: CRC Press, 2011), p. 324.
- ^ a b c d e Patel S, Gupta RS (December 2018). "Robust demarcation of fourteen different species groups within the genus Streptococcus based on genome-based phylogenies and molecular signatures". Infection, Genetics and Evolution. 66: 130–151. Bibcode:2018InfGE..66..130P. doi:10.1016/j.meegid.2018.09.020. PMID 30248475. S2CID 52813184.
- ^ Xu P, Alves JM, Kitten T, Brown A, Chen Z, Ozaki LS, et al. (April 2007). "Genome of the opportunistic pathogen Streptococcus sanguinis". Journal of Bacteriology. 189 (8): 3166–3175. doi:10.1128/JB.01808-06. PMC 1855836. PMID 17277061.
- ^ "Streptococcus". PATRIC. Blacksburg, VA: Virginia Bioinformatics Institute. Archived from the original on 2013-03-10.
- ^ a b Ferretti JJ, Ajdic D, McShan WM (May 2004). "Comparative genomics of streptococcal species". The Indian Journal of Medical Research. 119 (Suppl): 1–6. PMID 15232152.
- ^ McShan, W. Michael; Nguyen, Scott V. (2016), Ferretti, Joseph J.; Stevens, Dennis L.; Fischetti, Vincent A. (eds.), "The Bacteriophages of Streptococcus pyogenes", Streptococcus pyogenes: Basic Biology to Clinical Manifestations, Oklahoma City (OK): University of Oklahoma Health Sciences Center, PMID 26866212, retrieved 2024-02-07
- ^ McDonnell M, Ronda C, Tomasz A (1975) "Diplophage": a bacteriophage of Diplococcus pneumoniae. Virology 63:577–582
- ^ NCBI: Streptococcus phage Dp-1 (species)
- ^ Tiraby JG, Tiraby E, Fox MS (Dec 1975) Pneumococcal bacteriophages. Virology 68:566–569. doi:10.1016/0042-6822(75)90300-1. PMID 844
- ^ López R (September 2004). "Streptococcus pneumoniae and its bacteriophages: one long argument". International Microbiology. 7 (3): 163–171. PMID 15492930. PDF via web archive (9 Aug 2017)
- ^ Rubens López, Ernesto García: Recent trends on the molecular biology of pneumococcal capsules, lytic enzymes, and bacteriophage, Oxford Academic FEMS Microbiology Reviews. Volume 28, Issue 5. 1 Nov 2004, pp. 554—580, doi:10.1016/j.femsre.2004.05.002 (Free Fulltext)
- ^ Ronda C, López R, García E (November 1981). "Isolation and characterization of a new bacteriophage, Cp-1, infecting Streptococcus pneumoniae". Journal of Virology. 40 (2): 551–559. doi:10.1128/JVI.40.2.551-559.1981. PMC 256658. PMID 6275103.
- ^ Ouennane S, Leprohon P, Moineau S (2015). "Diverse virulent pneumophages infect Streptococcus mitis". PLOS ONE. 10 (2): e0118807. Bibcode:2015PLoSO..1018807O. doi:10.1371/journal.pone.0118807. PMC 4334900. PMID 25692983.
- ^ Johnsborg O, Eldholm V, Bjørnstad ML, Håvarstein LS (July 2008). "A predatory mechanism dramatically increases the efficiency of lateral gene transfer in Streptococcus pneumoniae and related commensal species". Molecular Microbiology. 69 (1): 245–253. doi:10.1111/j.1365-2958.2008.06288.x. PMID 18485065. S2CID 30923996.
- ^ Claverys JP, Håvarstein LS (March 2007). "Cannibalism and fratricide: mechanisms and raisons d'être". Nature Reviews. Microbiology. 5 (3): 219–229. doi:10.1038/nrmicro1613. PMID 17277796. S2CID 35433490.
- ^ Li G, Liang Z, Wang X, Yang Y, Shao Z, Li M, et al. (June 2016). "Addiction of Hypertransformable Pneumococcal Isolates to Natural Transformation for In Vivo Fitness and Virulence". Infection and Immunity. 84 (6): 1887–1901. doi:10.1128/IAI.00097-16. PMC 4907133. PMID 27068094.
- ^ Michod RE, Bernstein H, Nedelcu AM (May 2008). "Adaptive value of sex in microbial pathogens". Infection, Genetics and Evolution. 8 (3): 267–285. Bibcode:2008InfGE...8..267M. doi:10.1016/j.meegid.2008.01.002. PMID 18295550.
External links
[edit]- Centers for Disease Control and Prevention (CDC) (March 2000). "Adoption of perinatal group B streptococcal disease prevention recommendations by prenatal-care providers--Connecticut and Minnesota, 1998". MMWR. Morbidity and Mortality Weekly Report. 49 (11): 228–232. PMID 10763673.
- Nature-Inspired CRISPR Enzyme Discoveries Vastly Expand Genome Editing . On: SciTechDaily. June 16, 2020. Source: Media Lab, Massachusetts Institute of Technology.
- Streptococcus genomes and related information at PATRIC, a Bioinformatics Resource Center funded by NIAID
- The Canadian Strep B Foundation Archived 2013-05-02 at the Wayback Machine
- The UK Group B Strep Support charity
- Stuttering Streptococcal Infection Infection